Understanding Key elements of Molecular Biology
ContentTo understand Line Probe Assay (LPA), we need to understand key elements of Molecular Biology.
Molecular Biology is the study of living systems at the molecular level, especially DNA, RNA and proteins.
Cell
The cell is the fundamental unit of life and is the building block of all organisms. It has three main components:
- Cell membrane
- Nucleus
- Cytoplasm
Types of Cells
Cells are of two types- Prokaryotic and Eukaryotic and are distinguished by the size and types of internal organelles they contain.
- Prokaryotes are single cell organisms e.g., bacteria like Mycobacterium tuberculosis.
- Eukaryotes are either single celled or multicellular with membrane-bound organelles. All animals, plants, fungi and protists are eukaryotic cells.
DNA
- The DNA molecule is a nucleic acid containing genetic information and is made of two nucleotide strands bonded together.
- In prokaryotes, the DNA is present in the cytoplasm and in eukaryotes, it is present in the nucleus.
- The functional unit of the DNA is known as a gene.
RNA
- RNA is formed by the transcription of a DNA molecule
- RNA is also a nucleic acid but has a different nucleotide composition
- RNA translates to further form proteins, which are large molecules formed by one or more chains of amino acids
Table: Comparison between DNA and RNA
DNA RNA Double Strand Single Strand Transcribed to RNA Translated to Proteins Bases: adenine (A), cytosine (C), guanine (G), and thymine (T) Bases: adenine (A), cytosine (C), guanine (G), and uracil (U) Sugar Motif - Deoxy Ribose Sugar Motif - Ribose Stable Prone to Hydrolysis Resources
Kindly provide your valuable feedback on the page to the link provided HERE
Structure of DNA in a Prokaryotic Cell
Content- Deoxyribonucleic acid (DNA) is a molecule that contains genetic information.
- DNA molecules have two nucleotide strands that wind around one another to form double helix structure.
- Each nucleotide strand has a backbone made of alternating sugar (deoxyribose) and phosphate groups.
- Attached to each sugar is one of four nitrogenous bases: adenine (A), cytosine (C), guanine (G), and thymine (T)
- These bases occur in pairs, where in Adenine is always bonded to Thymine with a 2-H bonds and Cytosine bonded to Guanine with 3-H bonds.
- Prokaryotic cells do not have a defined nucleus hence the DNA is tightly coiled in the cytoplasm.
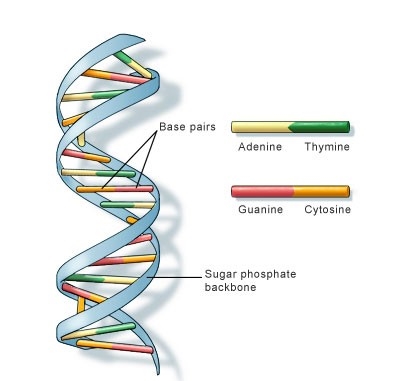
Figure: DNA double helix structure
Resources
Kindly provide your valuable feedback on the page to the link provided HERE
Central Dogma in the Function of DNA: Replication
ContentDeoxyribose Nucleic Acid (DNA) carry genetic information that is transmitted to new cells/off-springs.
The process by which cells maintain their genetic information and convert the genetic information encoded in DNA into gene products is called the Central Dogma.
The central dogma of DNA (Figure 1) are replication, transcription and translation.
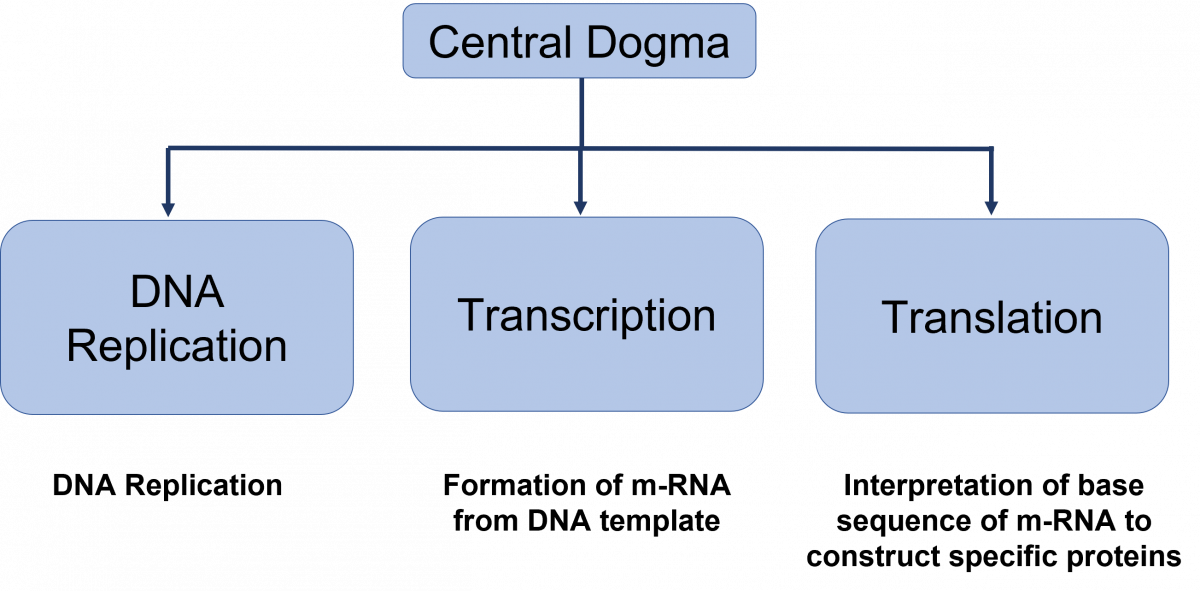
Figure: Central dogma of DNA functions

DNA Replication
DNA replication is the production of two identical replicas of DNA from one original DNA molecule.
Steps in DNA replication
- Helicase enzyme unzips coiled DNA to form a replication fork. The two separated strands act as templates for making new strands of DNA.
- Primase enzyme makes short primers to which DNA polymerase III enzyme binds and adds DNA nucleotides to create the leading strand during DNA replication.
- Primers are removed and replaced with DNA nucleotides by bacterial DNA polymerase I and DNA ligase seals the gaps between fragments.
- Okazaki fragments are short sequences of DNA synthesized discontinuously; linked by DNA ligase to create the lagging strand during DNA replication.
- Exonuclease enzyme removes primers and gaps are filled by DNA polymerase.
Resources
Kindly provide your valuable feedback on the page to the link provided HERE
Central Dogma in the Function of DNA: Transcription and Translation
ContentTranscription and translation together are responsible for Gene Expression.
Transcription
- The transcription process involves the formation of m-Ribo-Nucleic Acid (RNA) from DNA (Deoxyribose Nucleic Acid).
- One strand of DNA acts as a template, RNA polymerase associates with the DNA strand on the promoter region and transcription begins.
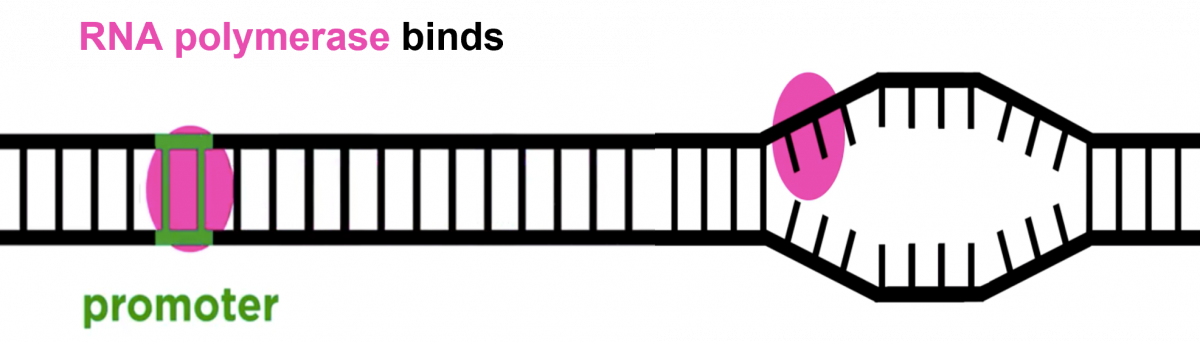
- RNA polymerase moves over the template DNA and adds complementary bases.
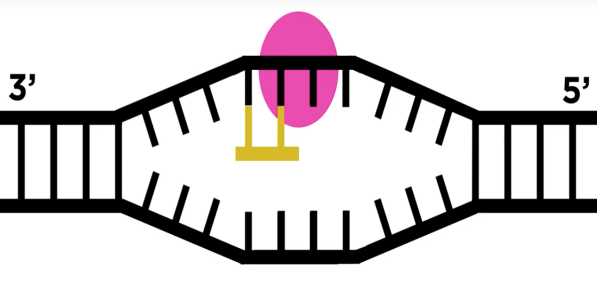
- RNA Polymerase stops transcription when the termination sequence is reached and releases the complete RNA chain, which moves to the cytoplasm.
Translation
- It is a process of synthesis of proteins from m-RNA.
- It starts when ribosomes attach to m-RNA.
- Bacterial ribosome has two subunits 50s and 30s, containing r-RNA and t-RNA.
- t-RNA is an adapter molecule - one side attaches to m-RNA, reading the triplet code and the other end attaches to a specific amino acid sequence.
- Once this assembly is formed, r-RNA catalyzes the process of attaching new amino acids, forming a chain.
The process of transcription and translation is shown in Figure 1
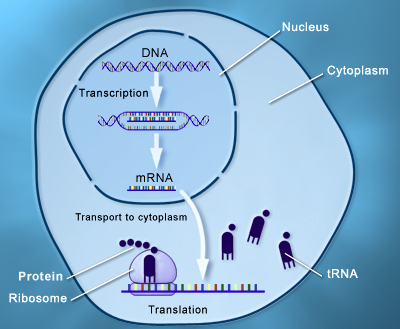
Figure 1: Process of transcription and translation under the central dogma of DNA function
Resources
Kindly provide your valuable feedback on the page to the link provided HERE
Molecular Basis of Genetic Code and Mutation
ContentThe genetic code is a triplet nucleotide sequence (codon) which encodes a specific amino acid during translation.
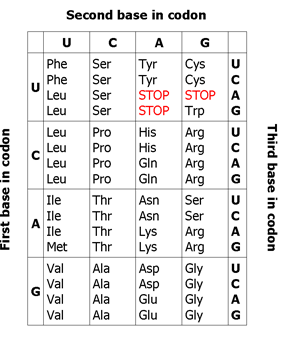
Figure 1: Genetic code
The three-letter codons consisting of four nucleotides found in mRNA (A, U, G, C) which produce a total of 64 different combinations. Of these 64 codons, 61 code for amino acids, the remaining three represent stop signals which trigger the end of protein synthesis (see Figure 1).
Properties of Genetic Code
- Universal: A particular codon will encode the same amino acid in all living beings
- Multiple codons: Different codons code for the same amino acid e.g. UUC and UUU code for Phe (Phenylalanine)
- Non-ambiguous: Each triplet specifies only a single amino acid
- Start codon (AUG): Codes for Met (Methionine) and marks the beginning of translation
- Stop codons (UAG, UAA, and UGA): Terminate protein synthesis as they do not code for any amino acid
Mutations
- Any alterations in the DNA sequence will result in an amino acid change e.g., GAA (Glutamic Acid) changes to GAC (Aspartic Acid)
- Changes in the amino acid sequence will alter the structure of the protein e.g., GAA (Glutamic Acid) changes to UAA (Stop codon)
- The structural change in the protein will alter its function e.g., for extracellular protein mutations in ACG (Cysteine) to TAT (Tyrosine) leads to loss of disulfide bond and protein function
- Silent mutations alter the DNA structure, but do not result in amino acid and functional changes e.g., GAA (Glutamic Acid) change to GAG (Glutamic Acid)
Resources
Kindly provide your valuable feedback on the page to the link provided HERE
Molecular Basis: Introduction to Polymerase Chain Reaction(PCR) Technology
ContentPolymerase Chain Reaction (PCR) is a laboratory technique to amplify Deoxyribose Nucleic Acid (DNA).
The PCR mix consists of:
- MgCl2 :1.5 - 6 mM
- Buffer (pH 8.3 - 8.8)
- DNA polymerase (Taq polymerase): 0.5 - 2.5 U
- Target DNA: <1µg
- Primers: Short DNA sequences to select the region to be amplified
During PCR, the temperature of the PCR mix is repeatedly raised and lowered to help the DNA polymerase enzyme replicate the target DNA sequence in the presence of the primer (Figure 1) and produce multiple copies in a few hours (Figure 2).
Steps in PCR:
- Denaturation of the DNA into single strands (94-95ºC)
- Annealing of primers to each strand for new strand synthesis (58-65ºC)
- Extension of the new DNA strands (72ºC)
The test is performed in a thermal cycler machine that maintains a different temperature required during the different PCR steps.
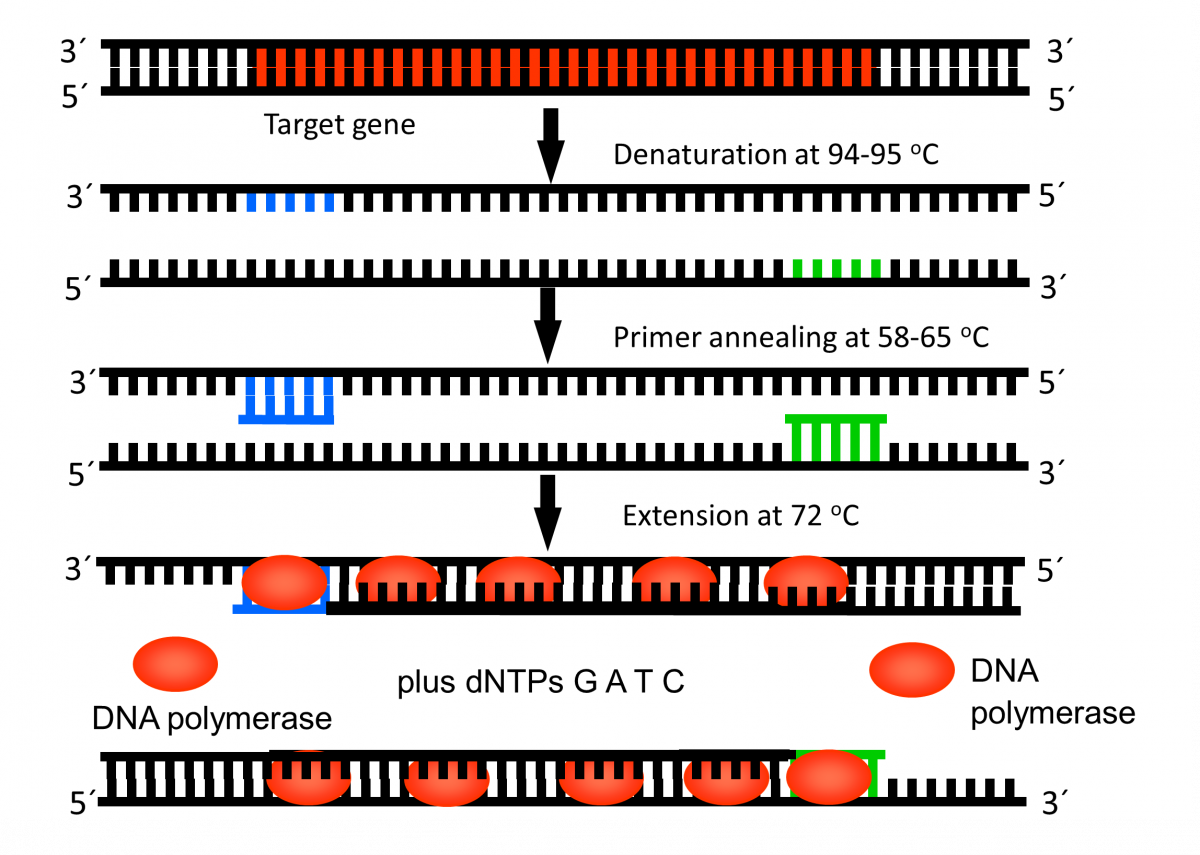
Figure 1: First Cycle in Thermal Cycler Machine

Figure 2: Exponential Amplification of Target Gene
Resources
Kindly provide your valuable feedback on the page to the link provided HERE
Molecular Mechanism of Multi-drug Resistance in Mycobacterium tuberculosis
ContentAnti-TB drugs and molecular mechanism of multi-drug resistance with regards to First Line- Line Probe Assay (FL-LPA) and Second Line- LPA (SL-LPA) is described here.
Rifampicin (RIF)
- It has a bactericidal effect on metabolically active M. tuberculosis and late sterilizing action on semi-dormant organisms undergoing short bursts of metabolic activity
- The mechanism of action of Rifampicin is the inhibition of RNA transcription in the mycobacterial cell by targeting DNA dependent RNA polymerase gene rpo B
- Resistance is due to mutations leading to a change in the structure of the beta subunit of Mycobacterium tuberculosis RNA polymerase
Isoniazid (INH)
- It has a potent early bactericidal action
- It is a pro-drug that requires activation by the mycobacterial enzyme, Catalase peroxidase gene katg
- INH resistant clinical isolates frequently lose their catalase peroxidase activity
INH may act on several targets within the mycobacterial cell, significant evidence supports the concept that it blocks the synthesis of cell wall mycolic acids. The major components of the envelope of M. tuberculosis.
Genes targeted in mycolic acid synthesis include:
- nadh-dependent enoyl acp synthase (encoded by inha)
- malonyl-coa acyl carrier protein (acp) transacylase (fabd)
- acetyl-coa carboxylase (accd6)
Other isoniazid target genes include peroxiredoxin alkyl hydroperoxide reductase subunit c (ahpc), ahpc-oxyrintergenic regulatory region; several efflux proteins encoded by iniabc and efpa
- Resistance is due to
- Mutations in katg, fabg1, oxyr- ahpc intergenic region, accd6 and efflux proteins
- Promoter region of inha, which leads to overexpression of isoniazid's target inha, requiring higher doses of the drug to achieve complete inhibition
Ethionamide (Eto)
- It has a bacteriostatic or bactericidal action, depending on the drug concentration in host
- It is a pro-drug, converted to active form by the bacterial monooxygenase EthA
- It targets mycolic acid synthesis
- Resistance is due to mutations in the inhA promoter
Pyrazinamide (Z)
- It has a bactericidal action; kills nonreplicating persistent Mycobacterium tuberculosis in macrophages
- It inhibits the synthesis of fatty acids; this disrupts the Mycobacterium tuberculosis cell membrane
- It is a pro-drug which must be activated by pyrazinamidase encoded by pncA, rpsA, and panD genes
- Resistance is due to mutation in pncA, rpsA, and panD; active drug effflux
Fluoroquinolones (Ofloxacin, Ofx; Levofloxacin, Lfx; Moxifloxacin, Mfx; Gatifloxacin, Gfx)
- Have a bactericidal action
- Inhibits gyrases encoded by gyrA and gyrB genes
- Prevent bacterial DNA synthesis
- High-level resistance requires multiple mutations in gyrA, or concurrent mutations in both gyrA and gyrB
Aminoglycosides (Kanamycin, Km; Amikacin, Am) /Polypeptides (Capreomycin, Cm)
- Have a bactericidal action
- Acts on ribosome rrs gene
- Prevents bacterial protein synthesis
- Resistance is due to mutation in rrs
- Low-level resistance to Kanamycin is associated with the promoter region of eis (enhanced intracellular survival protein) gene
Resources
Kindly provide your valuable feedback on the page to the link provided HERE
Fullscreen
