Interpretation of FL-LPA: Examples of Test Results Interpretation
Contentxamples of banding patterns for First Line - Line Probe Assay (FL - LPA) are shown in figure below.
Description
- All wild type bands display signal, mark as “+” against Wild Type (WT) column of the respective gene
- If at least one of the wild type bands is absent, mark as “-“ against WT column
- Similarly, “+” or “-” for Mutation (MUT) probes
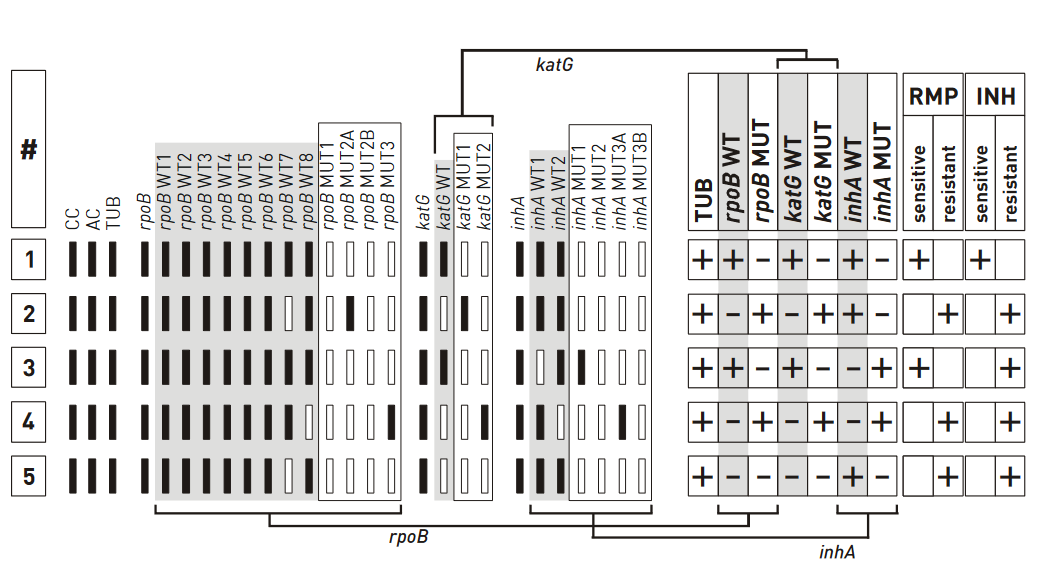
Figure: Banding patterns examples for Rifampicin and Isoniazid resistance; Source: GenoType MTBDR plus ver 2.0 kit, Instructions for Use.
Example #1
- The wild type banding pattern
- All wild type probes, but none of the mutation probes display a signal; hence, the evaluation chart shows “+”
- Three wild type columns and “–” in three mutation columns; hence “rifampicin (rmp) sensitive” and “Isoniazid (INH) sensitive” are marked “+”
Example #5
- One rpob and katg wild type probes missing; hence, the boxes for “rpob WT” and “katg WT” are marked with “–”
- No mutation probes developed; boxes are marked “–”
- inhA promoter region does not deviate from the wild type pattern, hence strain is evaluated as resistant to RMP and INH
Resources
Kindly provide your valuable feedback on the page to the link provided HERE
Evaluation of First-line Line Probe Assay Strip
ContentEach strip for First Line – Line Probe Assay (FL - LPA) has a total of 27 reaction zones (Figure below).
The Key Zones Include:
Conjugate Control (CC)
- Indicates the efficiency of conjugate binding and substrate reaction
Amplification Control (AC)
- Develops when control amplicon binds indication correct procedure was followed
- The absence of AC zone indicates an invalid test
M. tuberculosis Complex (TUB)
- Hybridizes with amplicons generated from Mycobacterium tuberculosis complex
- The absence of TUB zone indicates the absence of M. tuberculosis complex
Locus Controls (rpoB, katG, and inhA)
- Detect gene region specific for genes rpoB/ katg/ inhA
Wild-type Probes
- Comprise the resistance regions of respective genes (rpoB/ katg/ inhA)
- The absence of signal for at least one of the wild type probes indicates the resistance of the tested strain to the respective antibiotic
Mutation Probes
- Detect most common resistance-mediating mutations (rpoB/ katg/ inhA)
- Each pattern deviating from the wild-type pattern indicates resistance
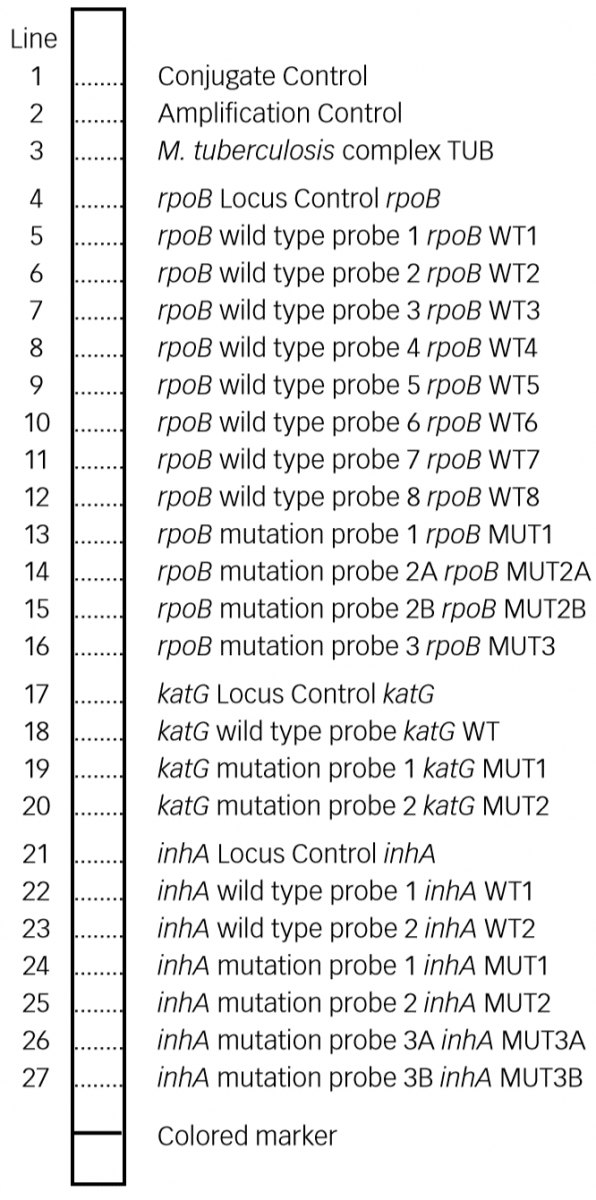

Figure: Key Zones of FL - LPA Evaluation Strip; Source: GenoType MTBDR plus ver 2.0 kit, Instructions for Use.
Evaluation of Assay Strip for FL - LPA Zones:
- The resistance status is determined and noted for each respective column.
- Distances between single probes on the strips may vary slightly.
- Not all bands of a strip have the same signal strength.
- The DNA strip should only be interpreted when:
- AC, CC and TUB bands are present
- Along with the presence of locus control bands for rpoB, katG and inhA
- Only those bands whose intensities are about as strong as, or stronger than that of the AC zone are to be considered.
- Bands whose intensities are about as strong as or stronger than that of the AC zone are considered.
Important: For an accurate evaluation, align provided template separately for each locus with the respective Locus Control band.
Resources
Kindly provide your valuable feedback on the page to the link provided HERE
Evaluation of Second-line Line Probe Assay Strip
ContentEach strip for Second Line - Line Probe Assay (SL - LPA) has a total of 27 reaction zones (Figure below).
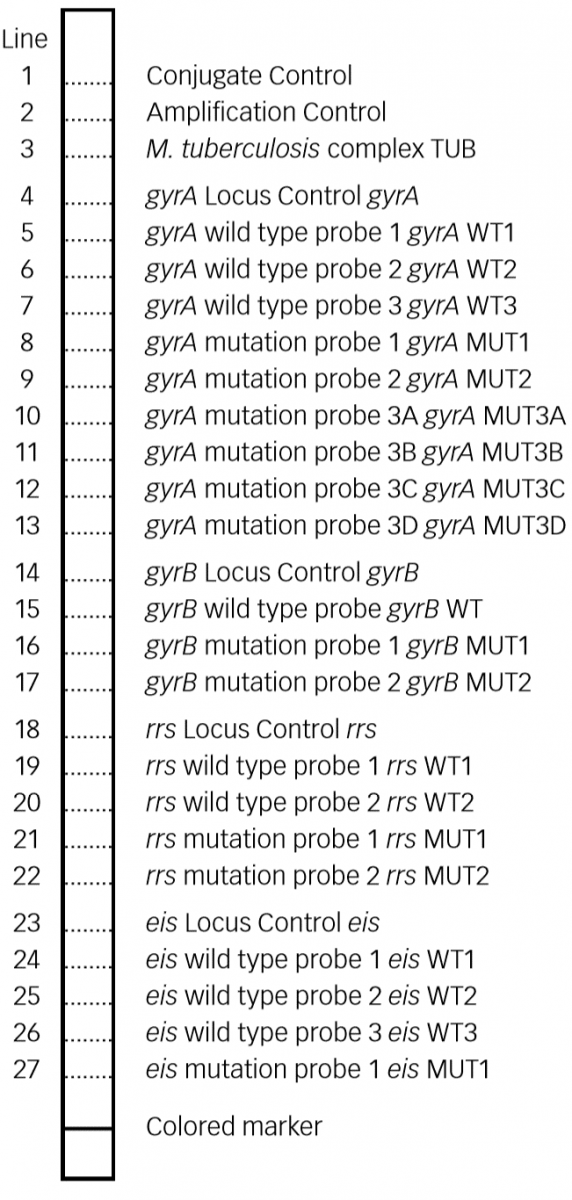

Figure: Key Zones of SL - LPA Evaluation Strip; Source: GenoType MTBDRsl VER 2.0 kit, Instructions for Use.
Evaluation of Assay Strip for SL- LPA Zones:
- Evaluation of Conjugate Control (CC), Amplification Control (AC), M. tuberculosis complex (TUB) are the same as described for First Line - LPA.
- Locus Controls (gyrA, gyrB, rrs, eis)
- Detect gene region specific (gyrA/ gyrB/ rrs/ eis)
- Wild type probes
- Comprise the resistance regions of respective genes (gyra/ gyrb/ rrs/ eis)
- Absence of signal for at least one of the wild-type probes indicates resistance of the tested strain to the respective antibiotic
- Mutation probes
- Detect most common resistance-mediating mutations (gyra/ gyrb/ rrs/ eis)
- Each pattern deviating from the wild type pattern indicates resistance
Important: For an accurate evaluation, align provided template separately for each locus with the respective Locus Control band.
Resources
Kindly provide your valuable feedback on the page to the link provided HERE
Interpretation of FL-LPA: Predicting TB Drug-resistance
ContentZone developed/ not developed (failing) in Wild Type (WT) and Mutation (MUT) probes are used to predict drugs resistance in First Line - Line Probe Assay (FL-LPA).
For Rifampicin
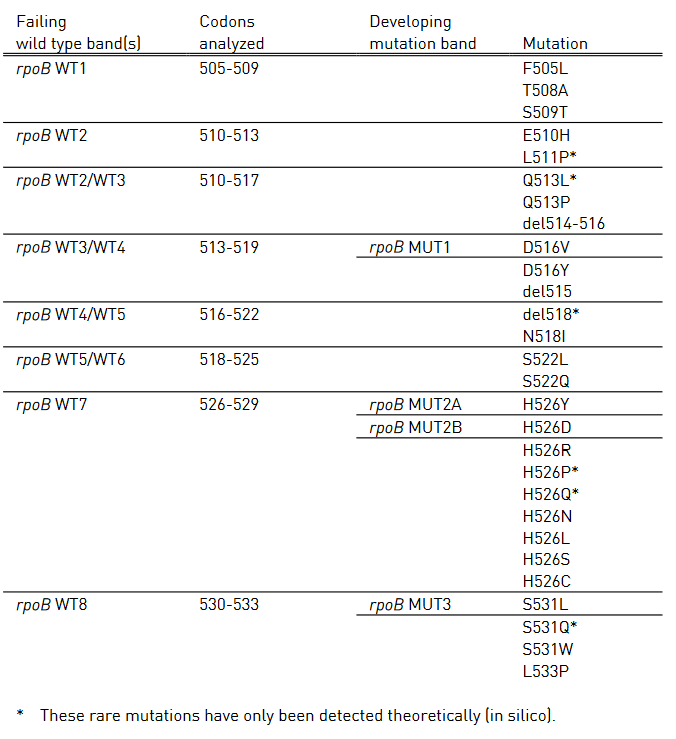
Table 1 shows mutations in rpoB gene and corresponding WT, MUT and responsible codons and mutations.
- rpoB WT1-8: rpoB wild type probes
- rpoB MUT1-3: rpoB mutation probes
Table 1: Mutations in the rpoB gene and corresponding wild type and mutation band; Source: GenoType MTBDR plus ver 2.0 kit, Instructions for Use.
For Isoniazid
Table 2 and Table 3 show mutations in katG gene and inhA promoter region, respectively and the corresponding WT and MUT bands.
- katG WT: katG wild type probe
- katG MUT 1-2: katG mutation probe
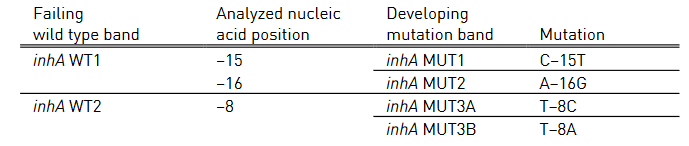
- inhA WT 1-2: inhA wild type probe
- inhA MUT 1-2, MUT 3A, MUT 3B: inhA mutation probe
Table 2: Mutations in katG gene and corresponding WT, MUT bands and responsible codons and mutations; Source: GenoType MTBDR plus ver 2.0 kit, Instructions for Use.
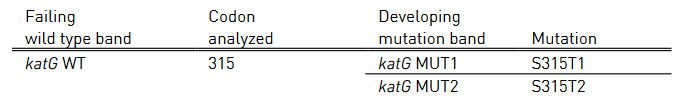
Table 3: Mutations in inhA promoter region and corresponding WT, MUT bands and responsible codons and mutations; Source: GenoType MTBDR plus ver 2.0 kit, Instructions for Use.
Resources
- GenoType MTBDR plus ver 2.0 kit, Instructions for Use.
- GLI Line Probe Assays for Drug-resistant Tuberculosis Detection.
Kindly provide your valuable feedback on the page to the link provided HERE
Interpretation of FL-LPA: Predicting Drug-resistance in Special Cases
ContentPredicting Drug Resistance in Special Cases
- When specimen tested contains a hetero resistant strain:
- Mutated as well as a wild-type sequence can be detected
- One of the mutation probes as well as corresponding wild type probe may stain positive on the respective strip
- Whether the respective resistance becomes phenotypically evident, depends on the ratio of mutated and nonmutated sequences
- The tested specimen contains more than one M. tuberculosis complex strain (due to mixed culture or contamination):
- One of these strains harbors a mutation, one of the mutation probes as well as the corresponding wild type probe may stain positive.
- Whether the respective resistance becomes phenotypically evident, depends on the ratio of the resistant and sensitive strain
- Due to mixed infection with M. tuberculosis complex strain and a nontuberculous mycobacterium:
- The TUB band may be missing
- If resistance pattern develops, presence of M. Tuberculosis complex must be suspected and repeat test
- All bands of a gene locus (including the Locus Control band) missing completely on a test strip:
- DNA concentration below the limit of detection or indicates the presence of interfering substances
- Repeat test
- If cultivated sample generates result with complete katG locus missing
- Indicates Isoniazid resistance
Resources
- GenoType MTBDR plus ver 2.0 kit, Instructions for Use.
- GLI Line Probe Assays for Drug-resistant Tuberculosis Detection.
Kindly provide your valuable feedback on the page to the link provided HERE
- When specimen tested contains a hetero resistant strain:
Interpretation of FL-LPA: Examples of Test Results Interpretation
Contentxamples of banding patterns for First Line - Line Probe Assay (FL - LPA) are shown in figure below.
Description
- All wild type bands display signal, mark as “+” against Wild Type (WT) column of the respective gene
- If at least one of the wild type bands is absent, mark as “-“ against WT column
- Similarly, “+” or “-” for Mutation (MUT) probes

Figure: Banding patterns examples for Rifampicin and Isoniazid resistance; Source: GenoType MTBDR plus ver 2.0 kit, Instructions for Use.
Example #1
- The wild type banding pattern
- All wild type probes, but none of the mutation probes display a signal; hence, the evaluation chart shows “+”
- Three wild type columns and “–” in three mutation columns; hence “rifampicin (rmp) sensitive” and “Isoniazid (INH) sensitive” are marked “+”
Example #5
- One rpob and katg wild type probes missing; hence, the boxes for “rpob WT” and “katg WT” are marked with “–”
- No mutation probes developed; boxes are marked “–”
- inhA promoter region does not deviate from the wild type pattern, hence strain is evaluated as resistant to RMP and INH
Resources
Kindly provide your valuable feedback on the page to the link provided HERE
Limitations of First-line Line Probe Assay [FL-LPA]
ContentFirst Line - Line Probe Assay (FL - LPA) can detect the mutations that are most frequently identified in resistant strains.
However, there are certain limitations of the test as follows:
- Some mutations that confer resistance are outside the regions covered by the test. Therefore, resistance cannot be completely excluded even in the presence of all wild type (WT) probes. Thus, in some cases, additional phenotypic Drug Sensitivity Testing (DST) may be necessary to provide a full assessment.
- Some mutations are identified specifically by mutation (MUT) probes, whereas others are only inferred by the absence of binding of the amplicons to WT probes. This lack of binding of a WT probe, without simultaneous binding of a MUT probe, is likely caused by the presence of a resistance mutation.
- Systematic errors are possible due to synonymous and non-synonymous mutations (e.g., phylogenetic mutations). Globally, this is rare (<1% of isolates), but these isolates can be frequent locally.
- LPA is less efficient than conventional culture-based methods in finding resistance in samples harbouring both drug-susceptible and resistant bacteria (i.e., hetero-resistance).
- More specifically, with LPA, it is possible to detect resistant bacteria with mutations detected by the MUT probes, if resistant bacteria represent at least 5% of the total population.
- However, resistant bacteria with mutations inferred by the absence of WT probes would probably be missed, if the resistant population is less than 95% of the total bacterial population.
Resources
Kindly provide your valuable feedback on the page to the link provided HERE
Reporting First-line Line Probe Assay [FL-LPA] Results
ContentResults of the First-line Line Probe Assay (FL-LPA) are entered in Nikshay and downloaded as the culture and drug susceptibility test (CDST) register.
Information on the patient, facility, reason for testing and test results are captured, and are shown in the table below.
DETAILS
TEST RESULTS
TestID
Visual_appearance_of_Sputum
DateOfTestUpdatedInNikshay
Sample
DateTested
TUB_Band
DateReported
RpoB_Locus_Control
TestStatus
RpoB_Wt1
Typeofspecimen
RpoB_Wt2
Dateofspecimencollection
RpoB_Wt3
StateName
RpoB_Wt4
DistrictName
RpoB_Wt5
TBUnit
RpoB_Wt6
PHINameLab
RpoB_Wt7
LabType
RpoB_Wt8
PatientId
RpoB_MUT1(D516V)
EpisodeID
RpoB_MUT2A(H526Y)
Name
RpoB_MUT2B(H526D)
Gender
RpoB_MUT3(S531L)
Age
KatG
PrimaryPhone
KatG WT1
Address
KatG MUT1
ResidentialState
KatG MUT2
ResidentialDistrict
InhA
TypeofCase
InhA WT1
ReasonforTesting
InhA WT2
TreatmentStatus
InhA MUT1
DiagnosisDate
InhA MUT2
TBTreatmentStartDate
InhA MUT3A
CurrentFacilityState
InhA MUT3B
CurrentFacilityDistrict
Final_Interpretation
CurrentFacilityTU
CurrentFacilityPHI
CurrentFacilityPHIType
ReasonforTesting1
PredominantSymptom
Predominantsymptom_duration
HistoryATT
No_of_HCP_visited_before_diagnosis_of_current_episode
Resources
- Guidelines for Programmatic Management of Drug-resistant TB, 2021
- Nikshay Reports CDST Test Register
Kindly provide your valuable feedback on the page to the link provided HERE
Reporting the Result of First-line Line Probe Assay [FL-LPA] in Special Cases
ContentReporting Drug-resistance in Special Cases
- When specimen tested contains hetero resistant strain:
- Strips contain both wild type and mutant probes for three genes known for resistance to Rifampicin and Isoniazid
- Bands to be interpreted as indicating resistance to a particular drug; should be reported as such
- Example of hetero resistant strain: rpoB WT 8 and rpoB MUT 3 bands are present
- Strips contain both wild type and mutant probes for three genes known for resistance to Rifampicin and Isoniazid
- The tested specimen contains more than one Mycobacterium tuberculosis complex strain (due to mixed culture/ non-tuberculous mycobacterium or contamination):
- If any strains harbor mutation, one mutation probe, as well as the corresponding wild type probe, may stain positive; ascertain the presence of TUB band
- Repeat test or phenotypic testing to confirm resistance
- Presence of non-tuberculous mycobacterium:
- TUB band missing; not be reported as M. tuberculosis
- TUB band absent but random banding patterns visible, eg: bands at rpoB WT 1, 4, 5 or 8; not be reported as M. tuberculosis
Resources
- GenoType MTBDR plus Ver 2.0 Kit, Instructions for Use.
- Line Probe Assays for Drug-resistant Tuberculosis Detection, GLI.
- Molecular Detection of Drug-resistant Tuberculosis by Line Probe Assay.
Kindly provide your valuable feedback on the page to the link provided HERE
- When specimen tested contains hetero resistant strain:
Interpretation of SL-LPA: Predicting Drug Resistance
ContentZone developed/ not developed (failed) in Wild Type (WT) and Mutation (MUT) probes are used to predict drugs resistance in Second Line - Line Probe Assay (SL - LPA).
For Fluoroquinolones (FLQ)
Tables 1 and 2 show mutations in gyrA, gyrB genes and corresponding WT, MUT and responsible codons and mutations:
- gyrA WT1-3: gyrA wild type probes; gyrA MUT1-2, 3A-3D: gyrA mutation probes
- gyrB WT: gyrB wild type probes; gyrB MUT1-2: gyrB mutation probes
Table 1: Mutations in gyrA gene and corresponding wild type and mutation band; Source: GenoType MTBDRsl Ver 2.0 Kit, Instructions for Use.
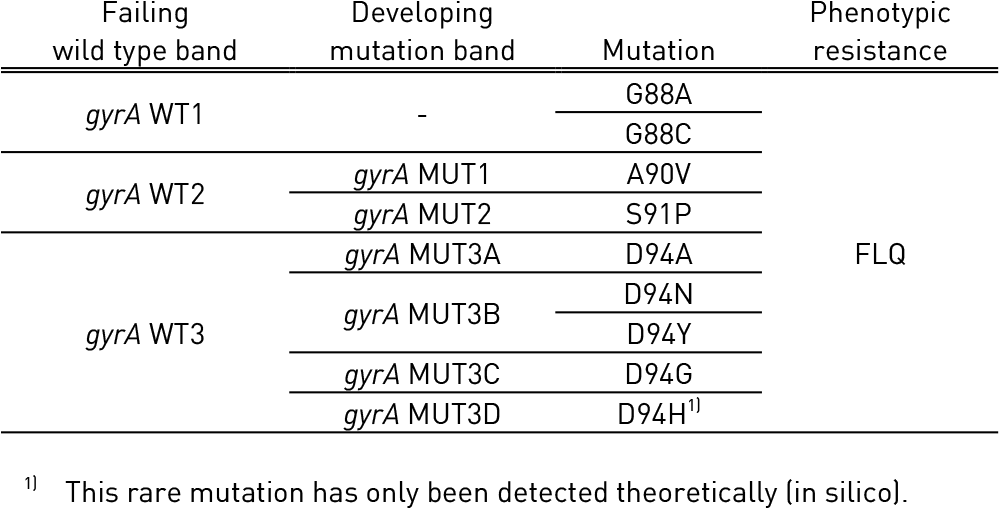
Table 2: Mutations in gyrB gene and corresponding wild type and mutation band; Source: GenoType MTBDRsl Ver 2.0 Kit, Instructions for Use.

For Second Line Injectable Drugs - Kanamycin (KAN), Amikacin (AMK), Capreomycin (CAP)
Tables 3 and Table 4 show mutations in rrs gene and eis gene respectively and corresponding WT and MUT bands:
- rrs WT 1-2: rrs wild type probe; rrs MUT 1-2: rrs mutation probe
- eis WT 1-3: eis wild type probe; eis MUT 1: eis mutation probe
Table 3: Mutations in rrs gene and corresponding WT, MUT bands and responsible codons and mutations; Source: GenoType MTBDRsl Ver 2.0 Kit, Instructions for Use.

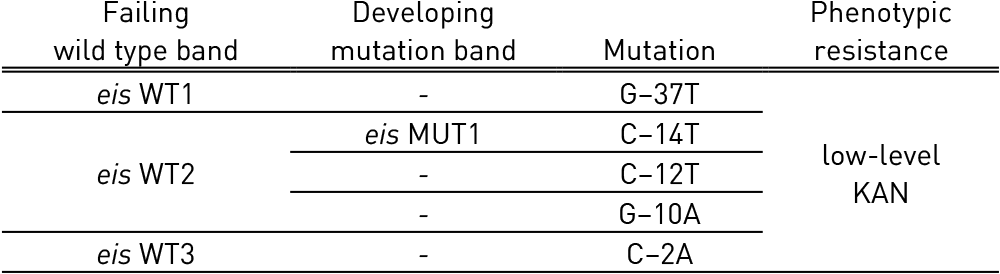
Table 4: Mutations in eis gene and corresponding WT, MUT bands and responsible codons and mutations; Source: GenoType MTBDRsl Ver 2.0 Kit, Instructions for Use.

Resources
- GenoType MTBDRsl Ver 2.0 Kit, Instructions for Use.
- Line Probe Assays for Drug-resistant Tuberculosis Detection, GLI.
Kindly provide your valuable feedback on the page to the link provided HERE
Interpretation of SL-LPA: Predicting Drug resistance in Special Cases
ContentNotes
- Bands with intensities similar or better than Amplification Control (AC) zone are considered.
- Not all bands of strip show same signal strength
Special Scenarios
- Both mutation probe and corresponding wild type probe are developed, represents a valid result
- Possible reasons:
- The tested specimen contains hetero-resistant strain
- The tested specimen contains more than one M. tuberculosis strain (mixed infection)
- Possible reasons:
- Resistance with wild type pattern
- Possible reasons:
- The tested specimen contains hetero-resistance strain; caused by mutation not covered by mutation probes
- The tested specimen contains the wild type and resistant strain (mixed infection); mutation not covered by mutation probes
- Possible reasons:
- Complete gene locus (all bands including Locus Control band) missing, invalid result
- Possible reason:
- DNA concentration below the limit of detection
- Possible reason:
Resources
Kindly provide your valuable feedback on the page to the link provided HERE
Interpretation of SL-LPA: Interpretation Examples
Contenthe examples of banding patterns for Second Line - Line Probe Assay (SL - LPA) are shown in the figure below.
Description
- All wild type bands display signal, mark as “+” against Wild Type (WT) column of the respective gene.
- If at least one of the wild type bands is absent, mark as “-“ against WT column. Similarly, “+” or “-” for Mutation probes (MUT).
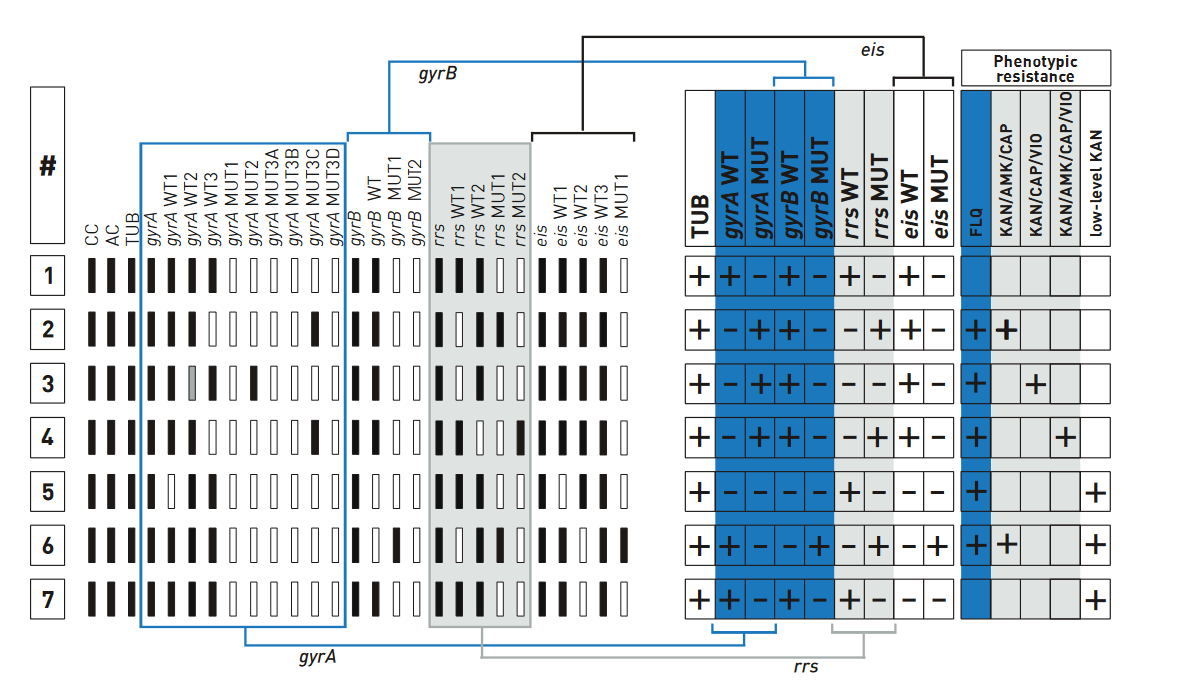
Figure: Examples for banding patterns for SL - LPA; Source: GenoType MTBDRsl Ver 2.0 Kit, Instructions for Use.
Example #1
- The wild type banding pattern.
- All wild type probes, but none of the mutation probes display a signal; hence, evaluation chart shows “+”.
- Four wild type columns and “–” in the four mutation columns; hence, no entry made in the fields of resistance columns.
Example #7
- gyrA locus and gyrB locus show wild type pattern; hence, evaluation chart shows “+” for fluoroquinolone (ofloxacin) resistance.
- rrs locus shows wild type pattern; hence, evaluation chart shows “+” for Kanamycin.
- eis wild type bands are present and “eis MUT” is absent; low-level Kanamycin resistance is assigned to the test.
Resources
Kindly provide your valuable feedback on the page to the link provided HERE
Limitations of Second-line Line Probe Assay [SL-LPA]
ContentThe limitations of Second Line Probe Assay (SL-LPA) are:
- It detects resistance due to mutation in gyrA, gyrB, rrs, and eis genes; resistances originating from mutations of other genes or gene regions as well as other drug resistance cannot be determined.
- Individual strains in mixed samples cannot be differentiated, which hampers the interpretation of the test.
- It cannot be used to monitor drug treatment, as DNA is detected from viable and non-viable bacteria.
- The results are qualitative, the intensity of bands does not give an estimate of bacterial load in the sample.
Resources
Kindly provide your valuable feedback on the page to the link provided HERE
Reporting Second Line-Line Probe Assay [SL-LPA] Results
ContentThe Results for Second Line Probe Assay (SL LPA) are entered in:
- Culture and Drug Susceptibility testing (CDST) register
- Nikshay-CDST module
- Request Form for examination of biological specimen for TB
- The fields entered are shown in Figure 1
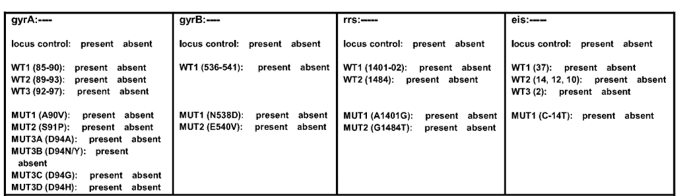
Figure 1: Result entry for Second Line LPA
Source: TOG Annexure- The final results are entered as:
- Select MTB Result: MTB Positive, MTB Negative
- Select Quinolone: Sensitive, Resistant, Indeterminate
- Select SLID (Second Line Injectable Drug): Sensitive, Resistant, Indeterminate
- Date result
- Date reported
- Reported by (name and signature)
Resources
- Challenge TB: LPA - Interpretation and reporting SL-LPA
- Guidelines for PMDT in India, 2021
- TOG Annexure
Kindly provide your valuable feedback on the page to the link provided HERE
Fullscreen